Taxonomy provides useful context for interpreting microbiome data, and taxonomy tables are often provided as output from microbiome sequencing pipelines or extracted from Bioconductor containers such as phyloseq. However, phylobar requires a phylo object as input. This is a more formal representation of a tree than a table of taxonomy assignments. To support the conversion, the package includes a helper function to convert such assignments into the required tree format. Since taxonomy tables lack a standardized representation, small discrepancies can produce invalid trees. This article collects examples from other vignettes of the common issues that can arise when constructing a taxonomy tree and goes into more depth about the strategies that were used to address them.
Checking that that the induced tree is correct
Before looking at potential issues, let’s review how to check whether
an input tree is valid. In the block below, we generate a random tree
using the rtree function from ape. We can confirm validity with the
checkValidPhylo function. If the output is TRUE, the tree
can be used for phylobar visualizations. If not, the function provides
hints on how to adjust the taxonomy table so that
taxonomy_to_tree produces a valid tree.
tree <- rtree(20)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 20
#> Found number of nodes: m = 19
#> Done.Introducing a missing root node
One common issue is the absence of a root node that links all descendant microbes. Many taxonomy tables implicitly assume the Bacteria kingdom and begin at the phylum level. In such cases, validity checks may fail with an error like the one below. This occurs because each phylum is treated as a separate tree, rather than being connected into a single tree with the Kingdom as the root.
data(atlas1006, package = "microbiome")
tree <- taxonomy_to_tree(tax_table(atlas1006))
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 128
#> Found number of nodes: m = 23
#> FATAL: nodes and tips should appear only once in the 2nd column of 'edge'
#> FATAL: the root node should not appear in the 2nd column of 'edge'
#> Done.This problem can be resolved by introducing a new “kingdom” column
from which all phyla descend. With this fix,
taxonomy_to_tree correctly builds edges between the first
and second columns. Re-running the validity check now produces a valid
tree object, which is used in the earlier Atlas vignette.
taxa <- tax_table(atlas1006)
taxa <- cbind(Kingdom = "Bacteria", taxa)
taxa <- phylobar::add_prefix(taxa)
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 130
#> Found number of nodes: m = 31
#> Done.Skipping over missing taxonomic assignments
Another common issue involves missing taxonomic assignments. phylobar can skip missing entries, but only if they are encoded as NA. If missing values are stored as character strings (e.g., “unclassified”), they are treated as valid categories. Since they often appear at multiple levels of resolution and under different parents, this breaks the tree structure.
download_zenodo("10.5281/zenodo.18791960", tempdir())
#> [zen4R][INFO] ZenodoRecord - Download in sequential mode
#> [zen4R][INFO] ZenodoRecord - Will download 1 file from record '18791960' (doi: '10.5281/zenodo.18791960') - total size: 76.2 KiB
#> [zen4R][INFO] Downloading file 'HFHS-data.rds' - size: 76.2 KiB
#> [zen4R][INFO] File downloaded at '/private/var/folders/mt/w0x_hxms30qch78n14v9zz0h0000gn/T/RtmpWbTXeT'.
#> [zen4R][INFO] ZenodoRecord - Verifying file integrity...
#> [zen4R][INFO] File 'HFHS-data.rds': integrity verified (md5sum: 3266a55a3d0e01db0f0c99c7cb7a8e06)
#> [zen4R][INFO] ZenodoRecord - End of download
HFHSdata <- readRDS(str_c(tempdir(), "/HFHS-data.rds"))
taxa <- HFHSdata$filtered_taxonomy
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 8
#> Found number of nodes: m = 287
#> FATAL: each tip must appear once in 'edge'
#> FATAL: all nodes should appear at least twice in 'edge'
#> MODERATE: some nodes are of degree 1 or less
#> FATAL: nodes and tips should appear only once in the 2nd column of 'edge'
#> Done.To avoid this, missing values should be explicitly coded as NA. This approach is illustrated in our high-fat, high-sugar diet vignette, where the taxonomy table is pre-processed to replace placeholder strings with NA. Here is the taxonomy before any correction. Notice that the NAs are not properly coded – we see the prefix coming from the taxonomic level, but no corresponding name.
head(taxa)
#> X1 X2 X3 X4 X5
#> OTU_13 OTU_13 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_21 OTU_21 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_7 OTU_7 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_29 OTU_29 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_30 OTU_30 k__Bacteria p__Firmicutes c__Clostridia o__Clostridiales
#> OTU_14 OTU_14 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> X6 X7 X8
#> OTU_13 f__S24-7 g__ s__
#> OTU_21 f__S24-7 g__ s__
#> OTU_7 f__S24-7 g__ s__
#> OTU_29 f__Rikenellaceae g__ s__
#> OTU_30 f__ g__ s__
#> OTU_14 f__S24-7 g__ s__We address this in the block below, checking for whether the name
ended with __ and replacing those with explicit NA
value.
taxa <- taxa |>
select(-X1, X1) |>
mutate(across(everything(), ~if_else(str_ends(., "_"), NA, .)))
head(taxa)
#> X2 X3 X4 X5
#> OTU_13 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_21 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_7 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_29 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> OTU_30 k__Bacteria p__Firmicutes c__Clostridia o__Clostridiales
#> OTU_14 k__Bacteria p__Bacteroidetes c__Bacteroidia o__Bacteroidales
#> X6 X7 X8 X1
#> OTU_13 f__S24-7 <NA> <NA> OTU_13
#> OTU_21 f__S24-7 <NA> <NA> OTU_21
#> OTU_7 f__S24-7 <NA> <NA> OTU_7
#> OTU_29 f__Rikenellaceae <NA> <NA> OTU_29
#> OTU_30 <NA> <NA> <NA> OTU_30
#> OTU_14 f__S24-7 <NA> <NA> OTU_14Once this transformation is made, the table can be used to construct a valid tree.
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 212
#> Found number of nodes: m = 80
#> Done.Avoiding duplicated names across different taxonomic levels
Another common problem arises when taxonomic names are not explicitly
distinguished across different levels of resolution. For example, in the
dietswap dataset, some phylum- and family-level assignments
share the same names. If uncorrected, taxonomy_to_tree
interprets these edges as loops, which produces an invalid tree. The
validity check will fail in this case. To see this, let’s first extract
the taxonomy table.
data("dietswap", package = "microbiome")
diet_temp <- subset_samples(dietswap, timepoint == 1)
diet <- subset_taxa(diet_temp, taxa_sums(diet_temp) > 0)
taxa <- tax_table(diet)Note the repeated phylum and family names. This causes the check to fail.
head(taxa)
#> Taxonomy Table: [6 taxa by 3 taxonomic ranks]:
#> Phylum Family
#> Actinomycetaceae "Actinobacteria" "Actinobacteria"
#> Aeromonas "Proteobacteria" "Proteobacteria"
#> Akkermansia "Verrucomicrobia" "Verrucomicrobia"
#> Alcaligenes faecalis et rel. "Proteobacteria" "Proteobacteria"
#> Allistipes et rel. "Bacteroidetes" "Bacteroidetes"
#> Anaerostipes caccae et rel. "Firmicutes" "Clostridium cluster XIVa"
#> Genus
#> Actinomycetaceae "Actinomycetaceae"
#> Aeromonas "Aeromonas"
#> Akkermansia "Akkermansia"
#> Alcaligenes faecalis et rel. "Alcaligenes faecalis et rel."
#> Allistipes et rel. "Allistipes et rel."
#> Anaerostipes caccae et rel. "Anaerostipes caccae et rel."
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 117
#> Found number of nodes: m = 22
#> FATAL: nodes and tips should appear only once in the 2nd column of 'edge'
#> FATAL: the root node should not appear in the 2nd column of 'edge'
#> Done.To address this, we can add a small prefix that encodes the taxonomic
rank of each assignment. The helper function add_prefix
supports this concatenation. At this stage, however, running the
validity check still raises an error. As in the Atlas example, the issue
is the absence of a root node connecting the different phyla. Adding a
“Kingdom” column resolves this by linking the trees together.
taxa <- phylobar::add_prefix(taxa)
taxa <- cbind(Kingdom = "k_Bacteria", taxa)
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 119
#> Found number of nodes: m = 30
#> Done.The issue of duplicated taxonomy names also appears in the Global Patterns dataset. For instance, several phylum- and class-level assignments share identical names.
data(GlobalPatterns, package = "phyloseq")
chlamydiae <- subset_taxa(GlobalPatterns, Phylum == "Chlamydiae")
taxa <- tax_table(chlamydiae)
head(taxa)
#> Taxonomy Table: [6 taxa by 7 taxonomic ranks]:
#> Kingdom Phylum Class Order Family
#> 100535 "Bacteria" "Chlamydiae" "Chlamydiae" "Chlamydiales" "Waddliaceae"
#> 2936 "Bacteria" "Chlamydiae" "Chlamydiae" "Chlamydiales" "Waddliaceae"
#> 24341 "Bacteria" "Chlamydiae" "Chlamydiae" "Chlamydiales" "Parachlamydiaceae"
#> 579085 "Bacteria" "Chlamydiae" "Chlamydiae" "Chlamydiales" "Parachlamydiaceae"
#> 547579 "Bacteria" "Chlamydiae" "Chlamydiae" "Chlamydiales" "Parachlamydiaceae"
#> 136933 "Bacteria" "Chlamydiae" "Chlamydiae" "Chlamydiales" "Parachlamydiaceae"
#> Genus Species
#> 100535 "Waddlia" NA
#> 2936 "Waddlia" "Waddliachondrophila"
#> 24341 NA NA
#> 579085 NA NA
#> 547579 "CandidatusProtochlamydia" NA
#> 136933 "CandidatusProtochlamydia" "CandidatusProtochlamydiaamoebophila"This duplication results in an invalid phylo.
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 5
#> Found number of nodes: m = 9
#> FATAL: all nodes should appear at least twice in 'edge'
#> FATAL: nodes and tips should appear only once in the 2nd column of 'edge'
#> Done.Again, we can resolve this using the add_prefix
function. This dataset raises one additional challenge: the ASV names
are stored only as row names, not as an explicit column in the taxonomy
table. Without this, we cannot reach the leaf nodes of the tree. The
solution is to introduce a new column containing the ASV
identifiers.
taxa <- data.frame(taxa)
taxa <- phylobar::add_prefix(taxa)
taxa$ASV <- rownames(taxa)
head(taxa)
#> Kingdom Phylum Class Order Family
#> 100535 K_Bacteria P_Chlamydiae C_Chlamydiae O_Chlamydiales F_Waddliaceae
#> 2936 K_Bacteria P_Chlamydiae C_Chlamydiae O_Chlamydiales F_Waddliaceae
#> 24341 K_Bacteria P_Chlamydiae C_Chlamydiae O_Chlamydiales F_Parachlamydiaceae
#> 579085 K_Bacteria P_Chlamydiae C_Chlamydiae O_Chlamydiales F_Parachlamydiaceae
#> 547579 K_Bacteria P_Chlamydiae C_Chlamydiae O_Chlamydiales F_Parachlamydiaceae
#> 136933 K_Bacteria P_Chlamydiae C_Chlamydiae O_Chlamydiales F_Parachlamydiaceae
#> Genus Species ASV
#> 100535 G_Waddlia <NA> 100535
#> 2936 G_Waddlia Waddliachondrophila 2936
#> 24341 <NA> <NA> 24341
#> 579085 <NA> <NA> 579085
#> 547579 G_CandidatusProtochlamydia <NA> 547579
#> 136933 G_CandidatusProtochlamydia CandidatusProtochlamydiaamoebophila 136933After this adjustment, checking the validity of the resulting tree still returns an error. In this case, the error can be safely ignored: it occurs when a tree contains nodes with more than two descendants. phylobar accommodates such multifurcations without issue.
tree <- taxonomy_to_tree(taxa)
checkValidPhylo(tree)
#> Starting checking the validity of tree...
#> Found number of tips: n = 21
#> Found number of nodes: m = 15
#> FATAL: all nodes should appear at least twice in 'edge'
#> Done.To verify, we can simply plot the phylo object directly before creating a phylobar visualization. You can check that this a static version of the same tree that we work with in the main Global Patterns vignette.
plot(tree)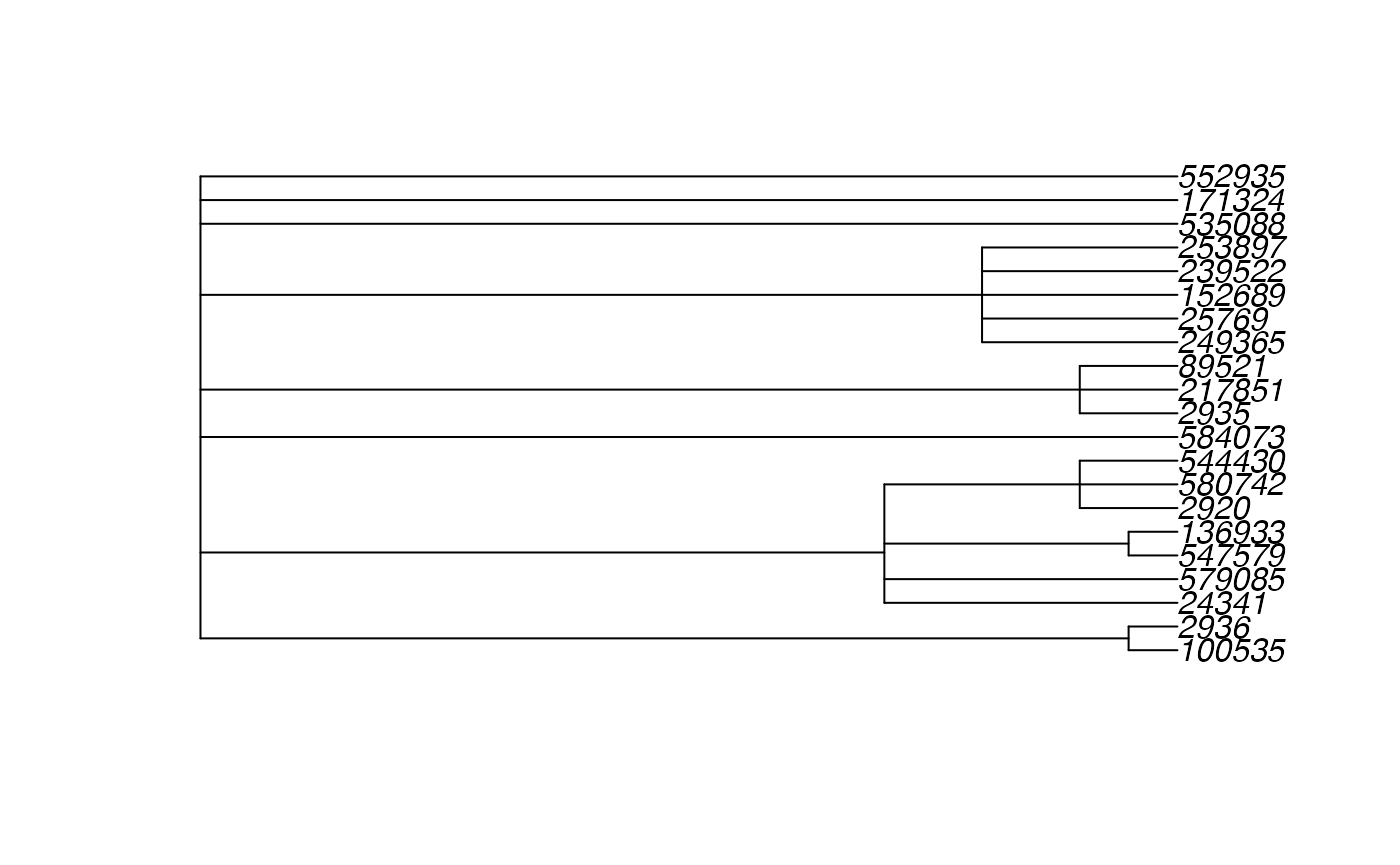
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: aarch64-apple-darwin23
#> Running under: macOS Sequoia 15.7.4
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.6/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.6/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> time zone: America/Chicago
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] zen4R_0.10.5 stringr_1.6.0 phyloseq_1.56.0 phylobar_0.99.12
#> [5] dplyr_1.2.1 ape_5.8-1 BiocStyle_2.40.0
#>
#> loaded via a namespace (and not attached):
#> [1] ade4_1.7-24 tidyselect_1.2.1
#> [3] farver_2.1.2 Biostrings_2.80.0
#> [5] S7_0.2.2 fastmap_1.2.0
#> [7] XML_3.99-0.23 digest_0.6.39
#> [9] lifecycle_1.0.5 cluster_2.1.8.2
#> [11] survival_3.8-6 magrittr_2.0.5
#> [13] compiler_4.6.0 rlang_1.2.0
#> [15] sass_0.4.10 tools_4.6.0
#> [17] utf8_1.2.6 igraph_2.3.1
#> [19] yaml_2.3.12 data.table_1.18.2.1
#> [21] knitr_1.51 phangorn_2.12.1
#> [23] S4Arrays_1.12.0 htmlwidgets_1.6.4
#> [25] curl_7.1.0 DelayedArray_0.38.1
#> [27] xml2_1.5.2 plyr_1.8.9
#> [29] RColorBrewer_1.1-3 abind_1.4-8
#> [31] withr_3.0.2 purrr_1.2.2
#> [33] BiocGenerics_0.58.0 desc_1.4.3
#> [35] grid_4.6.0 stats4_4.6.0
#> [37] multtest_2.68.0 biomformat_1.40.0
#> [39] ggplot2_4.0.3 scales_1.4.0
#> [41] iterators_1.0.14 MASS_7.3-65
#> [43] SummarizedExperiment_1.42.0 cli_3.6.6
#> [45] vegan_2.7-3 rmarkdown_2.31
#> [47] crayon_1.5.3 ragg_1.5.2
#> [49] generics_0.1.4 otel_0.2.0
#> [51] httr_1.4.8 reshape2_1.4.5
#> [53] cachem_1.1.0 splines_4.6.0
#> [55] parallel_4.6.0 BiocManager_1.30.27
#> [57] XVector_0.52.0 matrixStats_1.5.0
#> [59] vctrs_0.7.3 Matrix_1.7-5
#> [61] jsonlite_2.0.0 bookdown_0.46
#> [63] IRanges_2.46.0 S4Vectors_0.50.0
#> [65] systemfonts_1.3.2 foreach_1.5.2
#> [67] jquerylib_0.1.4 keyring_1.4.1
#> [69] glue_1.8.1 pkgdown_2.2.0
#> [71] codetools_0.2-20 stringi_1.8.7
#> [73] gtable_0.3.6 GenomicRanges_1.64.0
#> [75] quadprog_1.5-8 tibble_3.3.1
#> [77] pillar_1.11.1 htmltools_0.5.9
#> [79] Seqinfo_1.2.0 R6_2.6.1
#> [81] textshaping_1.0.5 evaluate_1.0.5
#> [83] lattice_0.22-9 Biobase_2.72.0
#> [85] bslib_0.10.0 Rcpp_1.1.1-1.1
#> [87] fastmatch_1.1-8 permute_0.9-10
#> [89] SparseArray_1.12.2 nlme_3.1-169
#> [91] mgcv_1.9-4 xfun_0.57
#> [93] fs_2.1.0 MatrixGenerics_1.24.0
#> [95] pkgconfig_2.0.3