Working with phyloseq and TreeSummarizedExperiment versions of the Global Patterns Dataset
Source:vignettes/global_patterns.Rmd
global_patterns.RmdThis vignette demonstrates how to create phylobar plots using either phyloseq or TreeSummarizedExperiment objects, two popular Bioconductor containers for microbiome data. We’ll work with the same dataset (GlobalPatterns) stored in both formats.
phyloseq Inputs
We will first study how to use phylobar on phyloseq objects. The block below sets the global pattern status and subsets to the Chlamydia subtree. We can extract the relevant sample composition using the otu_table accessor function.
data(GlobalPatterns, package = "phyloseq")
chlamydiae <- subset_taxa(GlobalPatterns, Phylum == "Chlamydiae")
x_phylo <- t(otu_table(chlamydiae))Phylogenetic Tree Tree
Our first approach will use the phylogenetic tree to support
hierarchical interaction. The tree can be extracted from the phyloseq
object using phy_tree. We first subsetted to only the
chlamydia subtree. There are now some samples without any observed
counts. Therefore, we will remove the samples from the view.
tree_phylo <- phy_tree(chlamydiae)
x_phylo <- x_phylo[rowSums(x_phylo) > 0, ]
x_phylo <- x_phylo / rowSums(x_phylo)We can now create our phylobar plot.
phylobar(x_phylo, tree_phylo, width = 900)Using Taxonomic Hierarchy
This creates the same type of visualization but using a taxonomic
hierarchy rather than a phylogeny. We need some problem-specific
preprocessing to deal with the fact that some ancestor and descendant
names look identical in this dataset. The add_prefix helper
ensures that the names are distinct between parents and children by
adding in the taxonomic rank to each name.
taxa <- tax_table(chlamydiae) |>
as.data.frame()
taxa$ASV <- rownames(taxa)
taxa <- phylobar::add_prefix(taxa)Now we can create a phylo tree object associated with the original taxonomy. Note that this is not a binary tree, since several taxonomic categories descended from a single node.
tax_tree <- taxonomy_to_tree(taxa)
plot(tax_tree)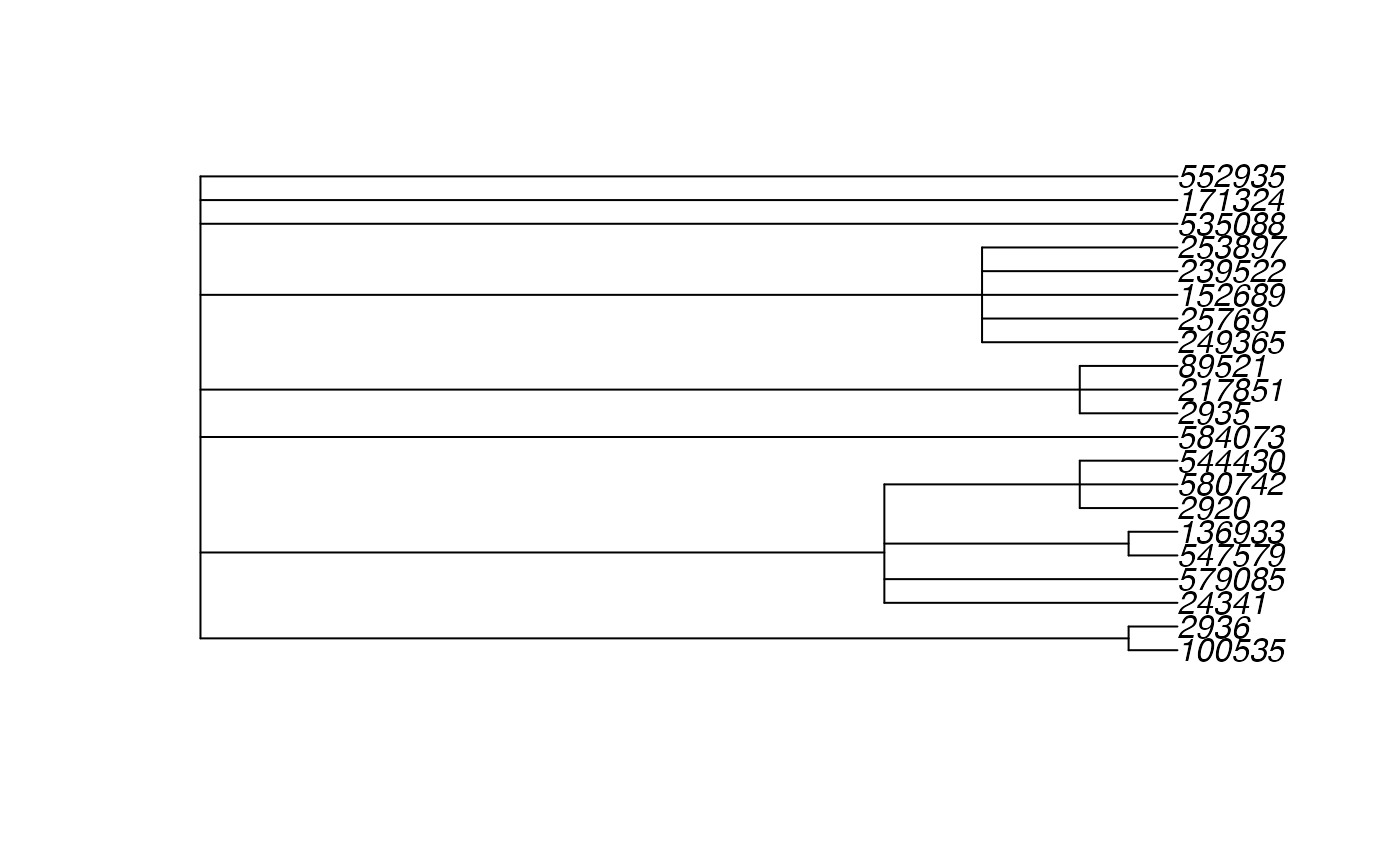
Given our new taxonomy and the earlier sample information, we can now create a phylobar plot.
phylobar(x_phylo, tax_tree, width = 900)TreeSummarizedExperiment Inputs
This example comes from the miaViz documentation and shows how to create a stacked bar tree from a TreeSummarizedExperiment. We’ll filter down to only those species that are present in at least one percent of all samples. In this example we’ll use a relative abundance transformation, so that we left with a composition of our plot rather than a more general stacked bar plot.
data(GlobalPatterns, package = "mia")
prev_species <- getPrevalent(GlobalPatterns, rank = "Species", detection = 0.01)
GlobalPatterns_tse <- GlobalPatterns[
rowData(GlobalPatterns)$Species %in% prev_species,
]
GlobalPatterns_tse <- transformAssay(
GlobalPatterns_tse,
method = "relabundance"
)Let’s keep only those taxes that are observed in at least one sample. We also need to filter the tree to reflect this reduced subset of taxa.
x_tse <- t(assay(GlobalPatterns_tse, "relabundance"))
x_tse <- x_tse[, colSums(x_tse) > 0]
tree_tse <- rowTree(GlobalPatterns_tse) |>
keep.tip(colnames(x_tse))With these inputs, we can generate the phylobar visualization. This view didn’t subset to the chlamydia class, which is why we have a larger tree here. But hopefully it’s clear that TreeSummarizedExperiment and phyloseq can essentially be used interchangeably when constructing the necessary inputs for these visualizations.
phylobar(x_tse, tree_tse, width = 900)
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: aarch64-apple-darwin23
#> Running under: macOS Sequoia 15.7.4
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.6/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.6/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> time zone: America/Chicago
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats4 stats graphics grDevices utils datasets methods
#> [8] base
#>
#> other attached packages:
#> [1] scater_1.40.0 scuttle_1.22.0
#> [3] miaViz_1.20.0 mia_1.20.0
#> [5] TreeSummarizedExperiment_2.20.0 Biostrings_2.80.0
#> [7] XVector_0.52.0 SingleCellExperiment_1.34.0
#> [9] MultiAssayExperiment_1.38.0 SummarizedExperiment_1.42.0
#> [11] Biobase_2.72.0 GenomicRanges_1.64.0
#> [13] Seqinfo_1.2.0 IRanges_2.46.0
#> [15] S4Vectors_0.50.0 BiocGenerics_0.58.0
#> [17] generics_0.1.4 MatrixGenerics_1.24.0
#> [19] matrixStats_1.5.0 ggraph_2.2.2
#> [21] ggplot2_4.0.3 phyloseq_1.56.0
#> [23] phylobar_0.99.12 ape_5.8-1
#> [25] BiocStyle_2.40.0
#>
#> loaded via a namespace (and not attached):
#> [1] RColorBrewer_1.1-3 jsonlite_2.0.0
#> [3] magrittr_2.0.5 ggbeeswarm_0.7.3
#> [5] farver_2.1.2 rmarkdown_2.31
#> [7] fs_2.1.0 ragg_1.5.2
#> [9] vctrs_0.7.3 multtest_2.68.0
#> [11] memoise_2.0.1 DelayedMatrixStats_1.34.0
#> [13] ggtree_4.2.0 htmltools_0.5.9
#> [15] S4Arrays_1.12.0 BiocNeighbors_2.6.0
#> [17] janeaustenr_1.0.0 gridGraphics_0.5-1
#> [19] SparseArray_1.12.2 sass_0.4.10
#> [21] bslib_0.10.0 tokenizers_0.3.0
#> [23] htmlwidgets_1.6.4 desc_1.4.3
#> [25] plyr_1.8.9 DECIPHER_3.8.0
#> [27] cachem_1.1.0 igraph_2.3.1
#> [29] lifecycle_1.0.5 iterators_1.0.14
#> [31] pkgconfig_2.0.3 rsvd_1.0.5
#> [33] Matrix_1.7-5 R6_2.6.1
#> [35] fastmap_1.2.0 tidytext_0.4.3
#> [37] aplot_0.2.9 digest_0.6.39
#> [39] ggnewscale_0.5.2 patchwork_1.3.2
#> [41] irlba_2.3.7 SnowballC_0.7.1
#> [43] textshaping_1.0.5 vegan_2.7-3
#> [45] beachmat_2.28.0 polyclip_1.10-7
#> [47] abind_1.4-8 mgcv_1.9-4
#> [49] compiler_4.6.0 fontquiver_0.2.1
#> [51] withr_3.0.2 S7_0.2.2
#> [53] BiocParallel_1.46.0 viridis_0.6.5
#> [55] DBI_1.3.0 ggforce_0.5.0
#> [57] MASS_7.3-65 rappdirs_0.3.4
#> [59] DelayedArray_0.38.1 bluster_1.22.0
#> [61] biomformat_1.40.0 permute_0.9-10
#> [63] tools_4.6.0 vipor_0.4.7
#> [65] otel_0.2.0 beeswarm_0.4.0
#> [67] glue_1.8.1 quadprog_1.5-8
#> [69] nlme_3.1-169 grid_4.6.0
#> [71] cluster_2.1.8.2 reshape2_1.4.5
#> [73] ade4_1.7-24 gtable_0.3.6
#> [75] tidyr_1.3.2 data.table_1.18.2.1
#> [77] BiocSingular_1.28.0 tidygraph_1.3.1
#> [79] ScaledMatrix_1.20.0 ggrepel_0.9.8
#> [81] foreach_1.5.2 pillar_1.11.1
#> [83] stringr_1.6.0 yulab.utils_0.2.4
#> [85] splines_4.6.0 dplyr_1.2.1
#> [87] tweenr_2.0.3 treeio_1.36.1
#> [89] lattice_0.22-9 survival_3.8-6
#> [91] tidyselect_1.2.1 DirichletMultinomial_1.54.0
#> [93] fontLiberation_0.1.0 knitr_1.51
#> [95] fontBitstreamVera_0.1.1 gridExtra_2.3
#> [97] bookdown_0.46 xfun_0.57
#> [99] graphlayouts_1.2.3 stringi_1.8.7
#> [101] ggfun_0.2.0 lazyeval_0.2.3
#> [103] yaml_2.3.12 evaluate_1.0.5
#> [105] codetools_0.2-20 gdtools_0.5.0
#> [107] tibble_3.3.1 BiocManager_1.30.27
#> [109] ggplotify_0.1.3 cli_3.6.6
#> [111] systemfonts_1.3.2 jquerylib_0.1.4
#> [113] Rcpp_1.1.1-1.1 parallel_4.6.0
#> [115] pkgdown_2.2.0 ecodive_2.2.6
#> [117] sparseMatrixStats_1.24.0 phangorn_2.12.1
#> [119] decontam_1.32.0 viridisLite_0.4.3
#> [121] tidytree_0.4.7 ggiraph_0.9.6
#> [123] scales_1.4.0 purrr_1.2.2
#> [125] crayon_1.5.3 rlang_1.2.0
#> [127] fastmatch_1.1-8